1 June 2026
Hurrah! Congratulations to Jenny Söderwind, who successfully defended her MSc thesis with the cool title: “Heat kills sex: Loss of sporulation and respiration in cold-tolerant Saccharomyces strains”.
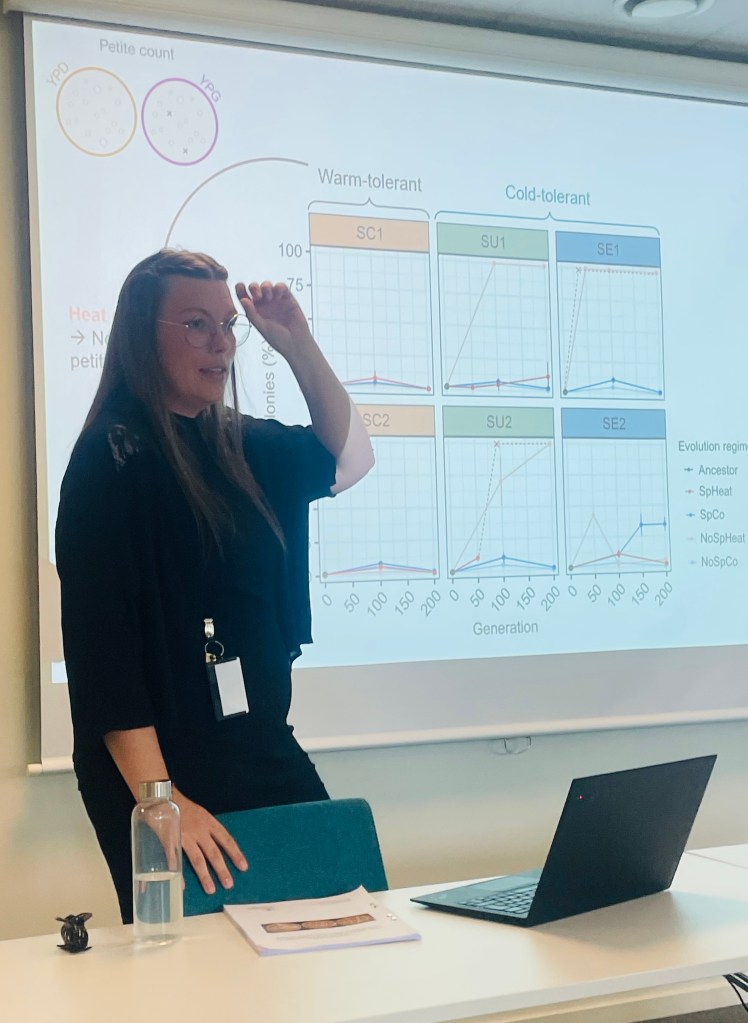
May 2026
Late announcement but better than never! We are welcoming a new lab member, Christian Oporto! Christian started his postdoc in our group in February (after graduating from his PhD at the University of Santiago, Chile with Francisco Cubillos). He now investigates the effect of temperature and other environmental stressors on yeast diversity and the evolution of pathogenicity. Christian is also passionate about fieldwork and loves hunting for interesting yeasts in the wild!

15 January 2026
Our new beer is ready! Yesterday, we got the chance to taste our new lager beer, brewed with a wild yeast strain from Patagonia. This is the result of a super fun collaboration (and lots of hard work) with Francisco Cubillos (U Santiago, Chile and iBio) and Andrés Furukawa at the Nils Oscar Brewery in Sweden. We will be telling the story behind the beer and our exciting journey to get it bottled at The Nobel Prize Museum in Stockholm on 13th February. See the invitation to the event here: https://www.nobelprizemuseum.se/en/event/the-chemistry-of-yeast-from-patagonia-to-local-beer/


10 November 2025
We’re welcoming a new PhD student to the lab! Rubie Min Lu, formerly in Jason Tsai‘s lab in Taiwan, has joined us in Stockholm. Welcome Rubie!

18 September 2025
We have a new paper out, now available in Molecular Ecology, in a Special Issue on Genomics of Speciation: “Reproductive isolation due to divergent ecological selection is accompanied by vast genomic instability in experimentally evolved yeast populations” https://doi.org/10.1111/mec.70110 In this paper, led by Devin Bendixsen, we evolved yeast for ~1,000 generations in divergent versus parallel environments. At regular time points during experimental evolution, we made crosses within and between environments, and show that adaptation to different “ecological niches” can generate partial reproductive isolation. Crosses between divergent-selected lines produced hybrids with reduced gamete viability and also striking genome instability (lots more SVs), while parallel-selected lines remained largely compatible. This links ecological selection to the emergence of “intrinsic” genetic barriers, and provides evidence that partial reproductive isolation can evolve due to adaptation to divergent environments, consistent with predictions of ecological speciation theory. We are particularly happy to see this out, as it has been in the making for a long time and was a large collaborative effort between postdocs, Phd students, and colleagues in Gothenburg!

11 July 2025
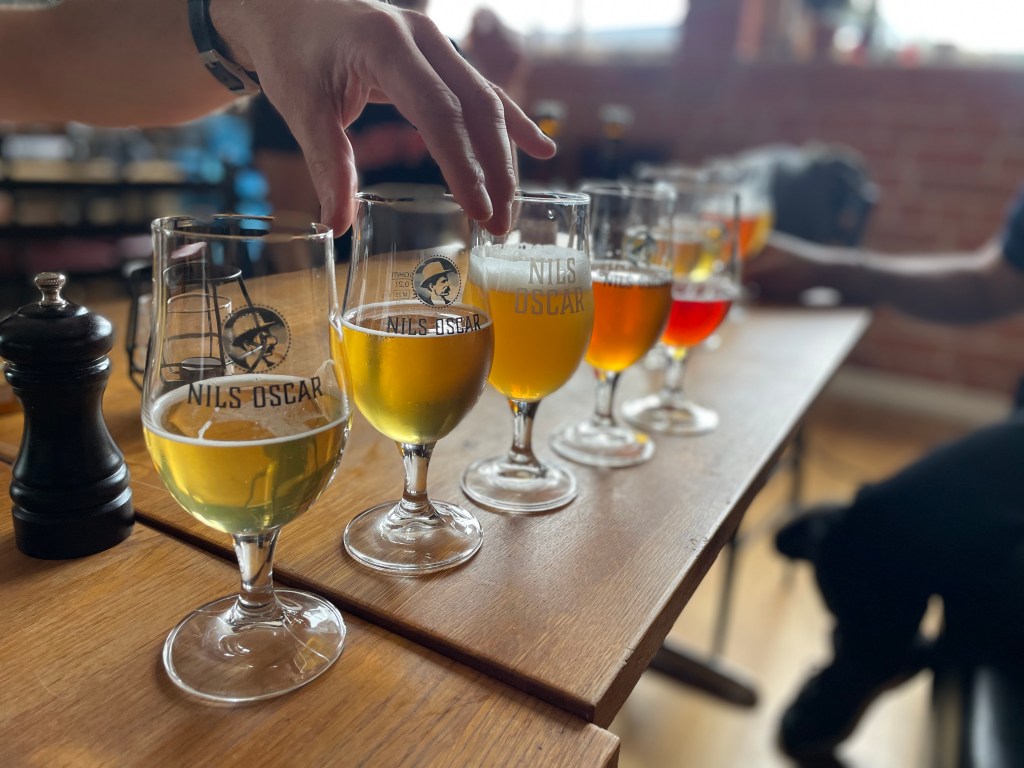
We’re still beaming from hosting illustrious guest for our workshop on “Yeast genomics and evolution” in Sweden! Thanks to Gilles Fischer (Sorbonne, France), Francisco Cubillos, and Pablo Villareal (U Santiago, Chile) for coming all the way to our remote European outpost! Thanks for sciencing, kayaking, walking, talking, and just generally making science as fun as it can be! We also visited our collaborators at the Nils Oscar Brewery in Nyköping (special thanks to skilled head brewmaster Andrés Furukawa), got to witness their brewing art in action, and got an amazing taste of their wide range of lagers and ales.




3 July 2025
Job alert! We have a postdoc position available! Join us and apply experimental evolution and comparative genomics in the model system yeast to test for parallel climate adaptation. You will develop a project that addresses fundamental questions in evolutionary biology. Specific aims are to 1) test for parallel climate adaptations in yeast from Europe and South America, 2) describe the genetic architecture of thermal traits, 3) test for the evolution of pathogenicity as a by-product of thermotolerance, and 4) use data-driven predictive modelling to identify key drivers of thermal adaptation. Please apply through this link: https://su.varbi.com/en/what:job/jobID:841228/ We are looking for candidates with a strong interest in evolutionary biology. Email me if you have any questions about the lab, work, and life in Sweden. Start date is negotiable but ideally in Oct 2025. Application deadline is 15 Aug 2025.
25 May 2025
Want to know more about the ecology of wild yeasts in Sweden, their association with oak trees, temperature, and rainfall? Read the latest by Chloé Haberkorn, Javier Pinto, and collaborators! https://doi.org/10.1111/1758-2229.70110. Celebratory chocolate cake in order!

20 May 2025
New paper out in PNAS! “The evolution of thermal performance curves in response to rising temperatures across the model genus yeast”. www.pnas.org/doi/10.1073/… Beautiful work by Jennifer Molinet, using experimental evolution to forecast the impact on global warming on biodiversity.
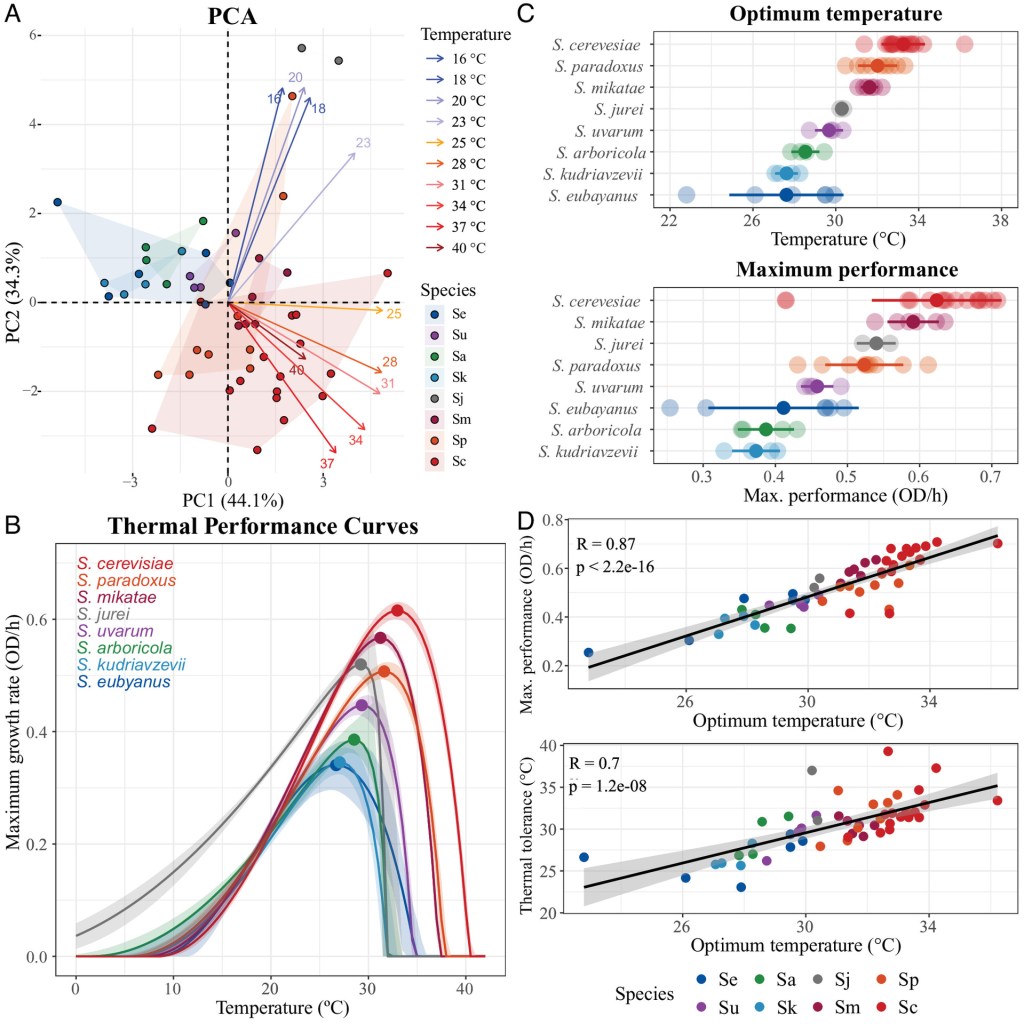
5 May 2025
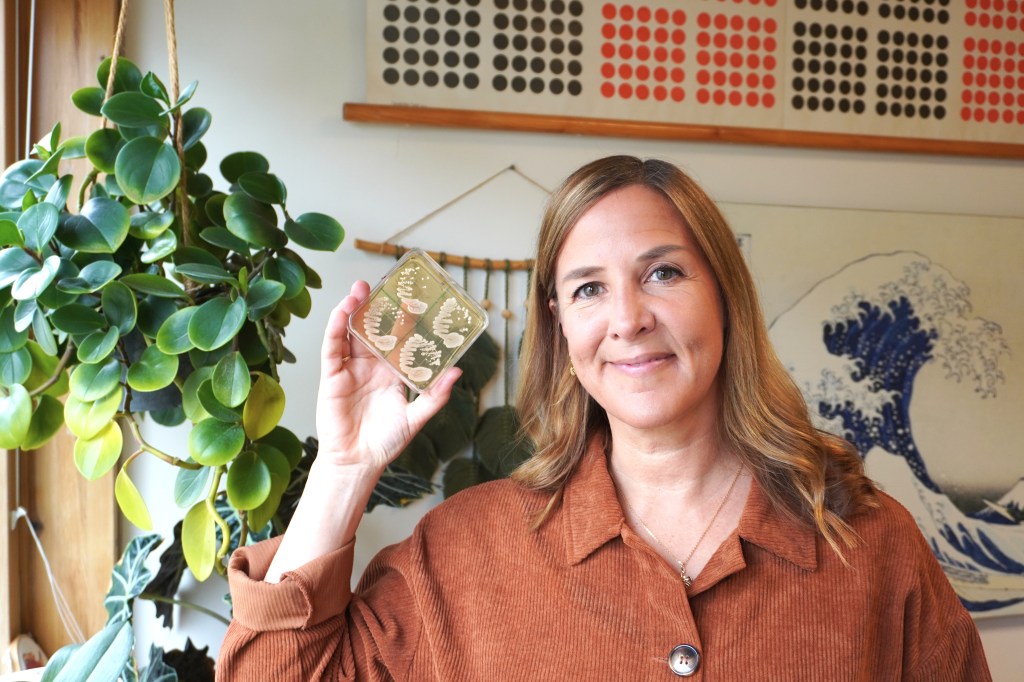
Stockholm University’s “Office for Research, Engagement and Innovation Services” came by and interviewed me about our collaboration with the beer industry. Link to article: https://www.su.se/english/news/how-rike-stelkens-turned-wild-yeast-into-a-new-kind-of-beer-1.819725

23 March 2025
We made our first beer! We used a yeast hybrid made from a strain collected in the forests of Patagonia in Chile and a commercial lager beer strain, and collaborated with a local brewery in Sweden to make two batches, one lager and one ale. Turned out pretty good! Now back to the lab for some refinements to get an even better flavour profile!

9 January 2025
Happy new year everybody! A quick little advertising here: Are you an undergraduate student interested in evolution and genomics? We (the Dept of Zoology and the Dept of Ecology, Environment and Plant Science at Stockholm University) are offering a brand new Master’s Programme in Evolutionary Genomics in the fall semester 2025! The application deadline is 15 January 2025 (really soon!). The programme offers a strong foundation for understanding the evolutionary processes that shape the genomic variation of populations. You will learn to use DNA sequencing technologies and bioinformatics tools to handle large amounts of data, understand what drives the evolution of populations and species, and address questions in biodiversity conservation. Check out the website and application criteria here: https://www.su.se/english/search-courses-and-programmes/nevgo-1.770335?open-collapse-boxes=programme-application

4 Dec 2024
Oooooh! We just received amazing grant news! More to be revealed soon, but to say this much: There will be both PhD and postdoc positions available in the Stelkens Lab!
1 Dec 2024
New paper alert: Carla Bautista, a PhD student with Christian Landry at the University Laval in Quebec in Canada, visited our lab 2 years ago. She just published a beautiful paper in Nature Communications that reveals how Haldane’s sieve can slow down adaptation in yeast hybrids! The paper is here: https://www.nature.com/articles/s41467-024-54105-4#citeas and Carla’s bluesky thread, where she explains her questions, methods and results, is here: https://bsky.app/profile/did:plc:tc5xl4aknyj66foiummjrypb/post/3lbzrisfhtc2j

15 Nov 2024
Congratulations to postdoc Chloé Haberkorn for winning a research grant from the Royal Physiographic Society of Lund, to investigate the phenotypic and transcriptomic responses to heat waves in different Saccharomyces species!
18 Sept 2024
The new fall semester is in full swing and we are welcoming Andrea Silva to the lab! Andrea (left in the picture) is a visiting PhD student from Tarragona in Spain, where she works on designing microbial consortia to improve wine fermentation! Pizza and wine to celebrate! Ok, some of us had a beer, too.

17 Sept 2024
I wrote about “messy” microbes and all the ways they do and not do not fit our traditional ideas of speciation (doi.org/10.1093/evolinnean/kzae023)

16 Sept 2024
We just had a short moment of fame on Swedish public television! Watch this clip, where Jennifer Molinet and I explain how we use wild yeast from Patagonia to make new types of Lager beer with fresh flavours and aromas, together with our collaborators from Chile!
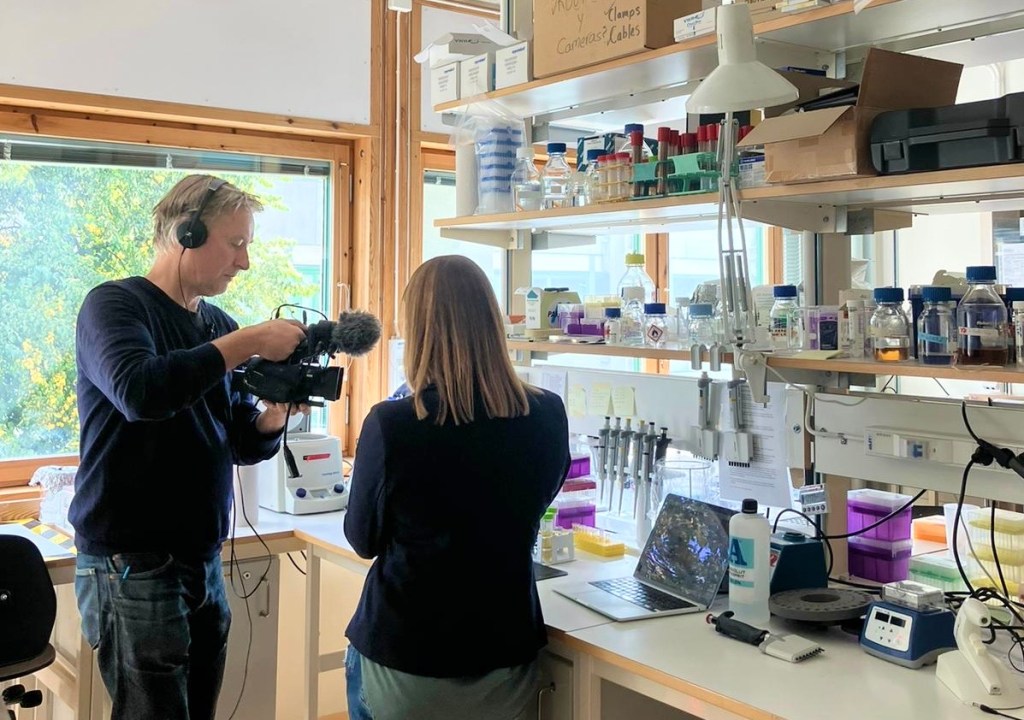

And here is an article in The Conversation that Jennifer wrote about our research (published in Plos Genetics, 2024): https://theconversation.com/weve-unlocked-exotic-new-beer-flavours-using-genetics-237266
1 June 2024
New paper out! Javier Pinto looked at two common estimates of population growth and fitness in microbiology – cell density (OD) and cell count (flow cyt) – across different species and strains of yeast. He found some surprises! First, there is a lot of variation in the slope of the cell density- cell count relationship, i.e. a given OD does not translate into the same number of cells across species (not even across strains within species). Generally, S. cerevisiae regressions were significantly steeper than the slopes found in other species (so, S. cerevisiae produces higher cell counts at lower OD). Interestingly, there is phylogenetic signal in the data, with more closely related strains producing more similar slopes. We thought this is perhaps explained by cell size, i.e. a given OD may result in fewer cell counts if cells are on average larger. But strangely, we found the opposite! At a given OD, strains with larger cells also produced higher cell counts. Puzzling! And quite unexpected. Whatever the reason, this means that if you are setting up competition experiments between strains that require equal starting population sizes or if you want to estimate fitness before and after experimental evolution, it’s worth calibrating and looking at strain specific parameters beforehand. Also, using just a single lab S. cerevisiae strain as a reference point to estimate the population fitness of wild, non-domesticated yeasts (most of which have much smaller cells) can be pretty misleading. We also provide a simple calculator in the paper to convert OD data into cell counts for each of the 23 strains we used here. Have a look! http://www.micropublication.org/journals/biology/micropub-biology-001215

5 May 2024
Look at that tiny, prickly oak bush growing on a wind swept, sandy beach in Greece. Would you have thought that it is covered in budding yeast (Saccharomyces cerevisiae)? Well, I didn’t! But here it is now, growing on our Petri dishes in the lab…
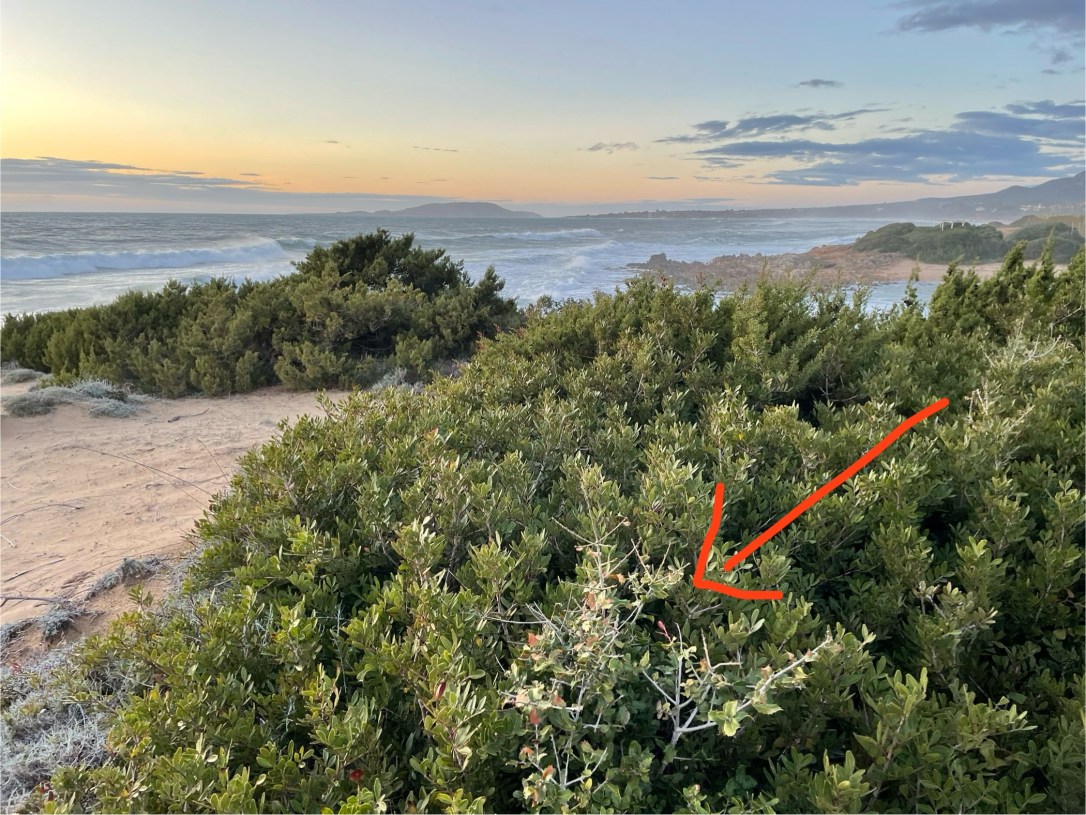

11 April 2024
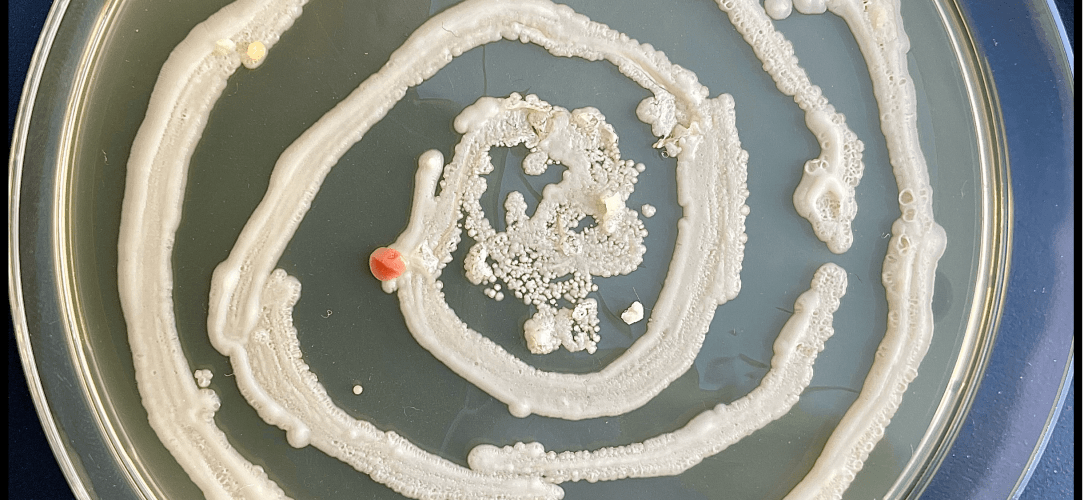
A sneaky and very gorgeous tiger eye contaminant in the lab (photo by Javier Pinto).
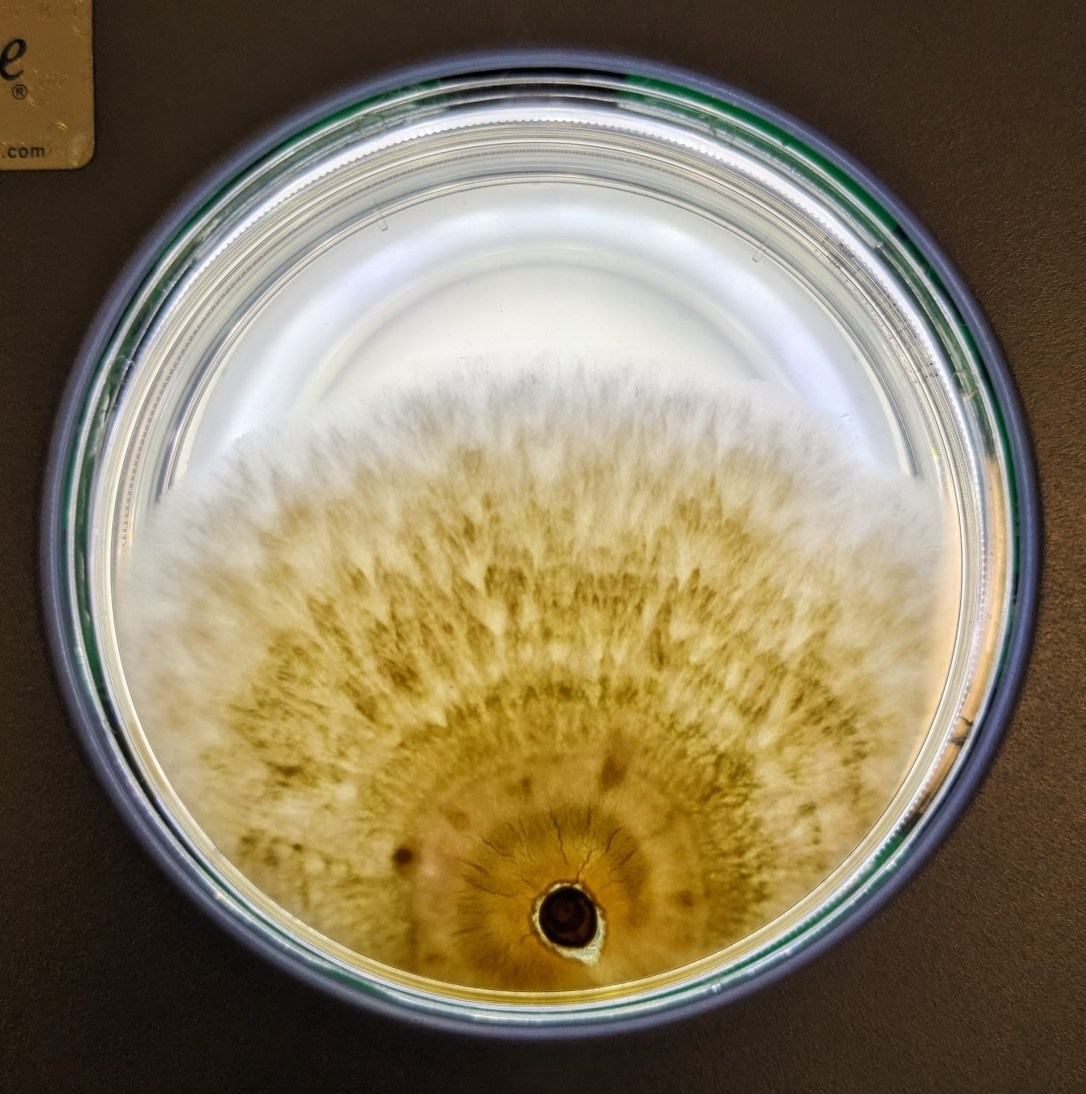

1 April 2024
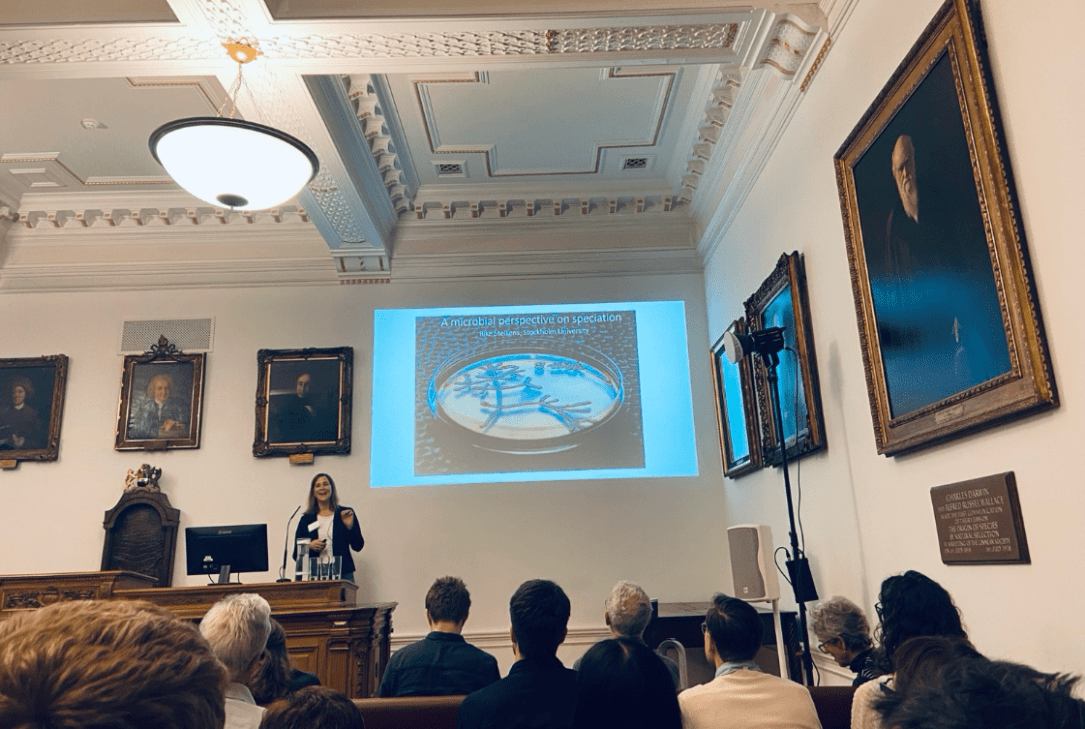
It was so much fun to give a “Microbial Perspective on Speciation” at the Linnean Society of London! Met old friends from my African cichlid fish days, got to know great new people, and saw inspiring talks. But the best treat of all was that the original portraits of Darwin and Wallace were with us in the room, watching over us! (photo by Domino Joyce).

15 March 2024
Congratulations to postdocs Javier Pinto and Chloé Haberkorn for receiving funding from the Bergström Foundation to conduct fieldwork in Värmland! Javier and Chloé will take yeast research back ‘into the wild’ and shed light on the biodiversity of Swedish forests using metabarcoding. Who knows, perhaps they will even find the elusive Saccharomyces eubayanus (one of the Lager beer yeast parents) hiding out in some cold place in the Swedish mountains!
13 March 2024
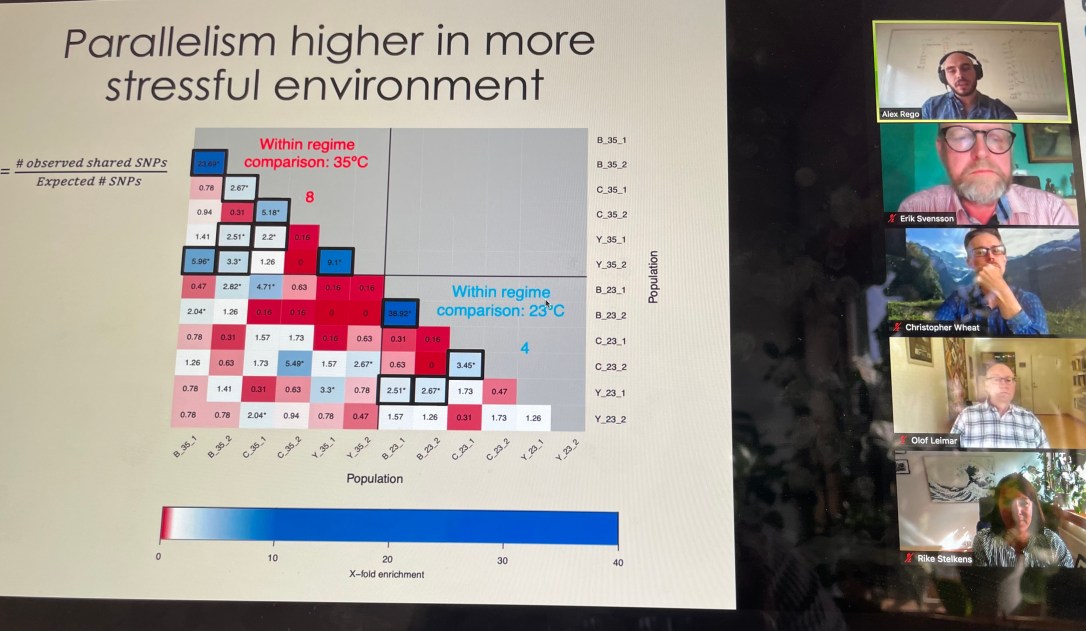
We have a new preprint out! Lead author Alex Rêgo found that “Adaptation to complex environments reveals pervasive trade-offs and genomic targets with large pleiotropic effects“. This work was inspired by the fact that most evolution experiments only consider a single stressor at a time. Few have addressed the question how adaptation proceeds in the presence of co-occurring stressors, which would be a much more realistic representation of natural environments. Alex adapted populations of yeast to a full-factorial combination of four stressors (heat, salt, starvation and an antifungal drug). He quantified the evolution of trade-offs between stressors, and the degree of genetic and phenotypic parallelism, and found some exciting results! Check it out: https://www.biorxiv.org/content/10.1101/2024.01.24.577006v2
5 February 2024
Here’s a gorgeous yeasty doodle for your enjoyment by Javier Pinto!

1 March 2024
If I was a yeast, I’d live on this oak tree by the waterfall in Greece.

30 January 2024
We have a new preprint out! “Wild Patagonian yeast improve the evolutionary potential of novel interspecific hybrid strains for Lager brewing” https://doi.org/10.1101/2024.01.29.577692 Lead author is the amazing Jennifer Molinet and this is a collaboration with our wonderful colleagues in Chile (Francisco Cubillos and Pablo Villareal among others). This paper has it all, wild yeast from Patagonia, hybrids, experimental evolution, whole genomes, de novo mutations, gene expression, gene copy number variation CRISPR, and of course, beer!

25 October 2023
We are welcoming 3rd year undergraduate student Elin to the lab! Elin will be working with Jenny on thermal adaptation of 8 Saccharomyces species from all around the world!

13 October 2023
To celebrate us as lab, to welcome new people and see ‘old’ ones go, we thought it was time to step up our Boule/Pétanque game with a glass of bubbly and some French delicatessen. Vive la Réboulution!



29 September 2023
Another student just graduated from the Stelkens lab! Alex Rêgo successfully defended his PhD thesis on the role of chance and determinism in evolution, using experimental evolution with small (yeast) and big bugs (seed beetles). A huge thank you to co-supervisor David Berger, to Dmitri Petrov, who was a stellar opponent for Alex thesis, and the amazing examining committee Tanja Slotte, Jonas Warringer and Martin Lind. Congratulations, Alex!
15 September 2023
And the excitement keeps going! This week, we are welcoming new postdoc Chloé Haberkorn in the lab! Not only is Chloé a brilliant evolutionary biologist and bioinformatician (check out her cool PhD work revealing a superlocus in the nightmarish bed bug that confers insecticide resistance), she’s also a fierce orienteering athlete in the French national team! Welcome Chloé, so happy you joined us!

8 September 2023
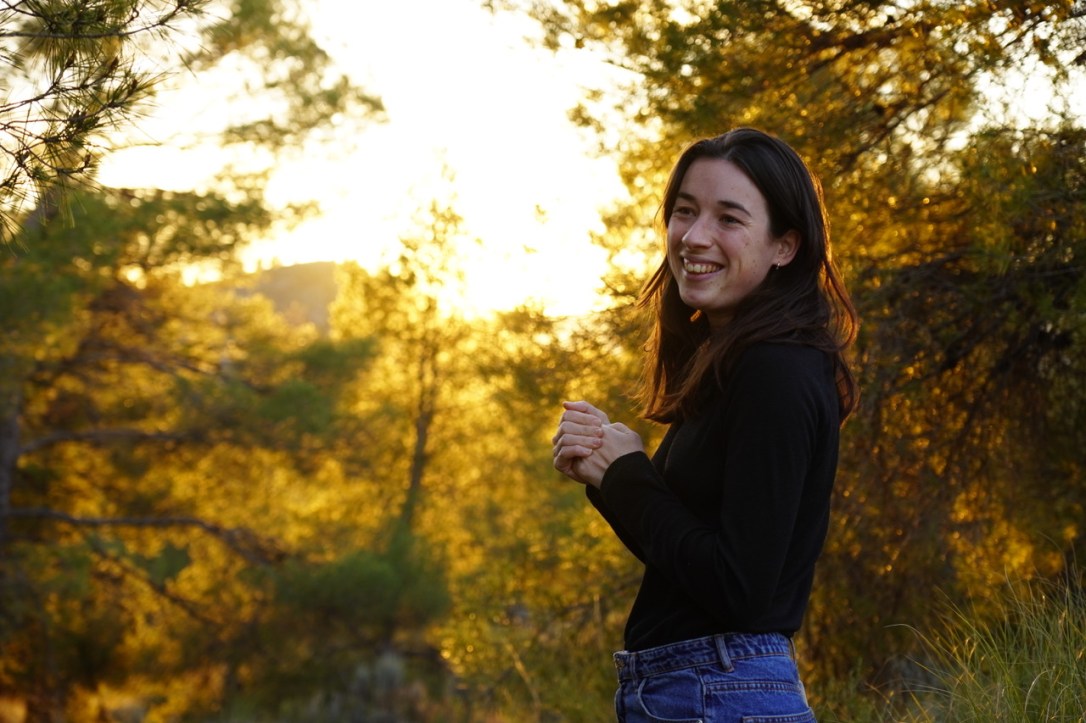
Exciting month in the Stelkens lab! First, a fabulous fieldtrip/mini-syposium at our zoological field station in Tovetorp, with our colleagues Francisco Cubillos and Pablo Villarreal from the University of Santiago (Chile), and Duncan Greig from UCL (UK). We were surrounded by beautiful ancient oaks, wild boar, a bat, and millions of mosquitos. Right now postdoc Javier Pinto is processing bark samples in the lab to see if we also find yeast (he thinks yes!).
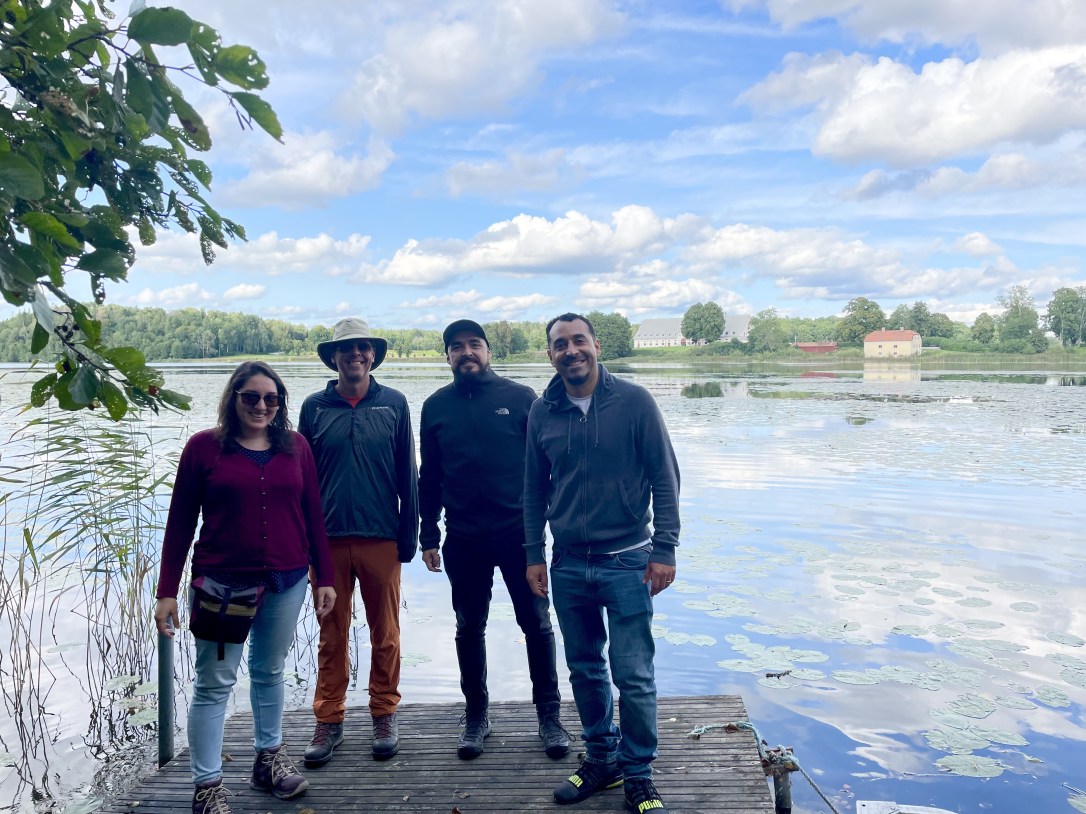







Next, PhD student Alex Rêgo completed his PhD thesis and, in old Swedish tradition, nailed it to a tree trunk outside the department for everyone to enjoy. Congratulations to Alex!!!
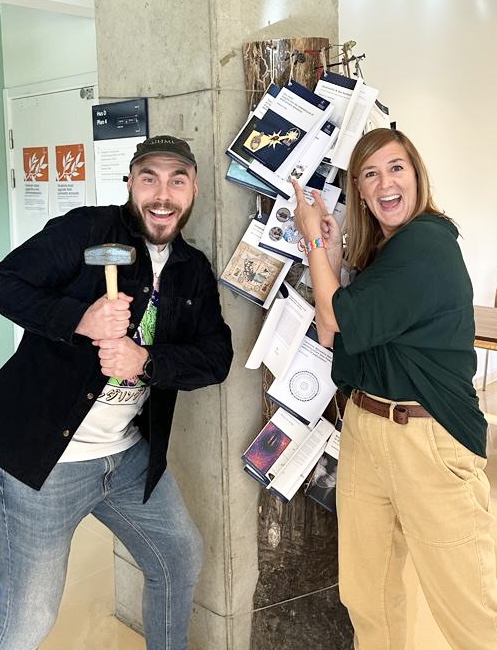

Finally, we had the great pleasure to host our friend Jasmine Ono, currently independent postdoctoral fellow at the University of Exeter (UK). Jasmine’s work on the evolution and genetic basis of reproductive isolation has been hugely influential for figuring out mechanisms of speciation (in yeast and beyond).

4 July 2023
Sampling Saccharomyces with Javier Pinto and our colleague Ayco Tack from an ancient oak tree on campus. This one had an ant colony living inside its trunk!

21 April 2023
A superwarm welcome to new postdoc Jennifer Molinet! Jennifer did it right, she left Santiago (where she worked in Fransisco Cubillo’s lab) at the end of the summer in Chile, just to join us in Stockholm at the beginning of our summer in Sweden! Jennifer comes with a huge backpack full of knowledge on yeast hybrids, experimental evolution, and molecular genetics, which she is already putting to good use in the lab. We’re very happy to have you with us, Jenny!
23 March 2023
We have a 2-year postdoc position available in “Genomics of Temperature Adaptation”. You will explore the genetic basis of adaptation to heat stress under relevant global warming scenarios, using long-term experimental evolution with eight yeast species from diverse ecological and geographic backgrounds, and functionally validate putative thermotolerance genes. Applications are due 30 April 2023. Start date in spring/summer 2023. Get in touch with Rike for any informal inquiries.
6 February 2023
New year, new visitors in the lab and exciting conferences to go to! Here are Luis-Miguel Chevin and Ken Thompson giving inspiring talks about the predictability of evolution in randomly fluctuating environments (Luis) and the role of ecology in hybrid incompatibilities (Ken). What a pleasure to host both of you!


And I had the great honour to present the lab’s work at the Gordon Research Conference on Speciation in Lucca (Tuscany)! The pool was cold but the conference was very warm and welcoming!


1 November 2022
Paper accepted in Molecular Biology and Evolution!! The dynamics of adaptation to stress from standing genetic variation and de novo mutations (open access). Congratulations to lead authors Lorena and Ciaran – oh-so-much sweat and tears went into this! These data from 1000 generations of experimental evolution of yeast in 4 environments with different fitness costs (stress levels) are the centerpiece of Ciaran’s PhD thesis, and when Lorena joined us a year ago, she went full steam ahead on the analysis of standing genetic variation vs. de novo mutations. Beautiful and interesting results, well done everyone involved!

14 October 2022
It’s a really BIG day! The first PhD student graduated from the Stelkens lab! Congratulations Dr. Ciaran Gilchrist to a beautiful PhD thesis on Hopeful Monsters: The Role of Hybrids in Adaptation. The party was wild, just as they say in the book!


8 September 2022
Exciting news! Postdoc Javier Pinto has just joined our lab. Javier just finished his PhD in Daniela Delneri’s lab at Manchester University, investigating genetic and epigenetic mechanisms of adaptation to cold temperatures in Saccharomyces. A super very warm welcome to Javier, so great to have you with us!

1 August 2022
Always tough to get back into the swing of things after the summer holidays but here we are. And luckily there are already new, motivated people waiting for us in the lab! Undergrad student Mario Walthert joins us today from Switzerland (formerly U Bern) to work with Ciaran and Alex on their experiments: The effect of hybridization in deteriorating environments, and adaptation to complex environments. Welcome Mario!

15 July 2022
Huge congratulations to Lorena Ament Velásquez who won a prestigious outgoing postdoctoral fellowship from the Swedish Research Council (Vetenskapsrådet)! Lorena will spend time in Pierre Gladieux‘s lab in Montpellier and Sven Saupe‘s lab in Bordeaux, and then return to Stockholm for her exciting research on “Self-nonself recognition systems as drivers of speciation in fungi” using the dung-loving fungus Podospora. So excited to see what Lorena finds and very happy she decided to stay with us for another 3 years!
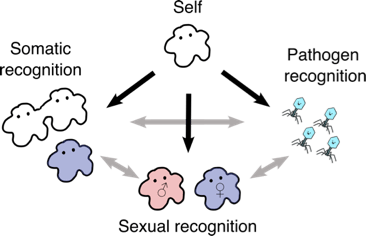
2 July 2022
We designed a survey on recent developments in speciation genetics and genomics! The survey primarily inquires about the species and systems that you study, the research approaches and techniques you use, and your opinions on the strengths and weaknesses of each. We hope it will highlight useful areas for future exploration and for technique/resource development. Please click on the following link to begin the survey, and feel free to share it with others. Thank you!
https://fresnostate.co1.qualtrics.com/jfe/form/SV_eXWcoGHyvJApv4q

27 June 2022
New paper alert! Heads up, if you are interested in lineage tracking with DNA barcodes. This is Shepherd, an accurate new clustering method to reduce noise and distinguish true from error sequences: Shepherd: accurate clustering for correcting DNA barcode errors. Shepherd enables higher resolution lineage tracking in your data and more accurate detection especially of small effect mutations. This is work with PhD student Nik Tavakolian in the lead, and born out of a fun and super inspiring collaboration with my friend and colleague Chun-Biu Li at the Math Department at Stockholm University. A Python implementation of Shepherd is freely available here. Thanks to the two kind, constructive and thorough reviewers at Bioinformatics who made this paper a lot better! Peer review at its best.
15 June 2022
I recruited two new lab assistants.




12 May 2022
Superfun night out with the lab, getting Ramen and Churros, celebrating that it is finally spring in Stockholm! We are also welcoming a new lab member, Jennifer Molinet (fourth from left). Jennifer is visiting us from Francisco Cubillo‘s lab in Chile. She is experimentally evolving S. cerevisiae x S. eubayanus hybrids – the first time we have that species combination in our lab! And *just* when we started digging into our Churros both Alex and Ciaran got the news that they’ll be giving talks at the ESEB in Prague in August! Whoop!

10 February 2022
News! I (Rike) am now Associate Editor at Evolution https://onlinelibrary.wiley.com/journal/15585646! Exited and proud to be given this role at one of my all time favourite journals that is working hard on making evolutionary biology a more diverse and inclusive place (check out https://diversifyeeb.com). Excited to see your submissions! And if you are an early career researcher, let me know, I’ll do my best to send a paper review your way.
20 January 2022
We are absolutely delighted to have PhD student Carla Bautista Rodriguez from Christian Landry’s lab (Université Laval, CA) with us for a three month lab visit! Carla is a brilliant young scientist working on yeast hybrid genomics. Welcome Carla!

5 January 2022
Happy new year! The end of 2021 was eventful for the lab, we published two papers on yeast hybrids with Devin Bendixsen in the lead! Both papers are fully open access.
In the first paper, Patterns of Genomic Instability in Interspecific Yeast Hybrids With Diverse Ancestries (with David Peris Navarro as collaborator) we looked at patterns of genomic instability in interspecific yeast hybrids with different ancestries, evolutionary histories, and ecologies (wine, beer, whiskey, baking, fruit, olives, clinical, lab strains). We were wondering if there are any patterns in hybrid genome composition across genetic and ecological backgrounds that would predict hybrid fitness. Devin tackled this with some smart comparative genomics. Using shared k-mer analysis, we found that more genetically distant crosses produce more similar hybrid genomes. We think this may be because stronger negative epistasis at larger genomic divergence puts constraints on hybridization outcomes. Using read depth analysis, we found that a whooping 44% of hybrid genomes carry at least one aneuploidy, affecting smaller chromosomes more often. Interestingly also, hybrids with more equal, more balanced content from both species were more likely to carry aneuploidies. We think this is because older hybrid genomes had more time for postzygotic processes (LOH, gene conversion, backcrossing), resulting in more stable genomes. All this is really fun, and shows there are definitely some shared patterns across ancestries and across ecology that can help us understand what role hybridization plays in adaptation, and allows a peek into the genetic architecture of hybrid fitness. But of course there is lots we do not know. E.g. what about structural variation, copy number variation, translocations and inversions? Do hybrids have more than their parents? Long read seq will soon tell us more we hope!

In the second paper, Saccharomyces yeast hybrids on the rise, we compiled all genomic resources available of the eight Saccharomyces species and their interspecific hybrids. To illustrate the degree and potential causes of reproductive isolation, we reanalyzed published data on hybrid spore viabilities across the entire genus and tested for the role of genetic, geographic, and ecological divergence within and between species (28 cross types and 371 independent crosses, see figure below). Hybrid viability generally decreased with parental genetic distance likely due to antirecombination and negative epistasis, but notable exceptions exist, highlighting the importance of strain-specific structural variation and ploidy differences. Surprisingly, the viability of crosses within species varied widely, from near reproductive isolation to near-perfect viability. Geographic and ecological origins of the parents predicted cross viability only to an extent.
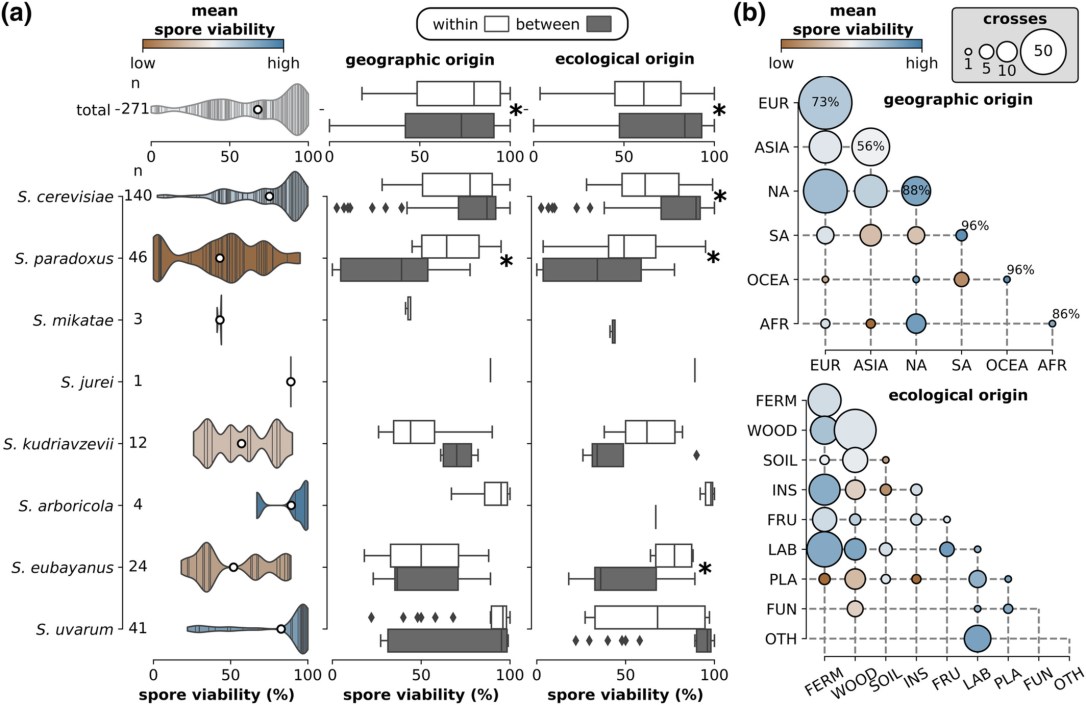
25 November 2021
It was such a pleasure to host Carla Bautista from Christian Landry’s lab at Université Laval (Québec) for a zoom talk and chat about yeast, hybrids, the universe, life on other planets, and adaptation to extreme conditions! So inspirational, thanks Carla! Looking forward to hosting you in our lab in the spring!

8 September 2021
A very happy WELCOME to our newest lab member and postdoc Lorena Ament Velásquez! Lorena came to us from Hanna Johannesson‘s lab at Uppsala University where she did her PhD on the population dynamics of fungal meiotic drive elements in Podospora. We are very happy to have you on board and are looking forward to sharing the power of yeast genetics with you. Use it wisely! 🙂
20 August 2021
At last, we meet again! Superfun Lab Party on a beautiful late Swedish summer day, with stand-up-paddling, pizza, s’mores, and delicious cakes. Looking forward to a fun fall and winter semester with these good people!






16 August 2021
Good bye and farewell to Dragan, who is off to new postdoc adventures in Switzerland! Dragan wasn’t with us for a long time, but for a REAL TIME!

15 June 2021
Congratulations to PhD student Alex Rêgo for successfully defending his half time thesis Theoretical and empirical approaches to infer determinism in evolution! Many thanks to the examining committee Erik Svensson, Chris Wheat and Olof Leimar for a great discussion of Alex’ work. Looking forward to continuing our conversation offline when we can all meet in person again.
14 May 2021
We have a new paper out in The American Naturalist! https://www.journals.uchicago.edu/doi/10.1086/715356 We show that “Hybridization outcomes have strong genomic and environmental contingencies” using seven interspecific yeast hybrid crosses. This was an awesome, concerted effort between former and current postdocs Claire, Zebin and Devin. Team work at its best! It has been a dream of mine since I was an undergrad to publish in this journal. Huge shout out to the editors and reviewers who made this paper so much better.
Have you ever wondered why some hybrids are fit and fertile and others doomed? What features of the parents’ phenotypes and genomes can help predict this? And does hybrid fitness depend on the environment they grow in at all? We use quantitative genetics theory and growth along environmental stress gradients to answer these and other questions. Turns out (as so often), it’s complicated! The number of transgressive (i.e. extreme) F2 hybrids varied significantly among environments, cross backgrounds, and their interactions, suggesting that hybridization outcomes are highly context-specific even when hybrid crosses are made from the same two parents. But it would be so useful to predict hybrid fitness better in the future, right? Think agricultural breeding and food production under extreme climate conditions. Transgressive hybrid plants and animals may be better at surviving droughts or flooding! Well, we’ll keep working on this to see if we can get past the idiosyncrasies caused by specific genetic backgrounds. We’ll focus more on the targets of selection in hybrid genomes, the genetic architecture of these traits, and their interactions with the environment.
This paper’s celebratory cake: lemon drizzle

6 April 2021
It is official! We are looking for another postdoc to join our Yeast Evolution and Genomics lab at Stockholm University in Sweden. Applicants will have a strong interest in evolutionary processes, especially in adaptation mechanisms, speciation, and conservation genetics, and be familiar with the principles of quantitative and population genetics. Ideally, the candidate has experience in experimental evolution, molecular genetics/genomics, and bioinformatics. Prior training in experimentation with Saccharomyces yeast and strong quantitative skills are desirable. Applicants should be well-organized, self-motivated, good communicators, and happy to work in an international team. Applicants must hold a PhD in evolutionary biology or a similar subject. The position (100% research) is for 1 year with the possibility for extension.
Members of our lab are currently working on a range of topics including the effects of hybridization on rates and mechanisms of adaptation, the genetic basis of hybrid fitness, adaptation to environmental stress, epistasis, fitness landscapes, speciation mechanisms, transposable element evolution, and phylogenomics. We use both experimental and theoretical approaches. While the project the applicant will work on is flexible and depends on personal preferences and ideas, the applicant is encouraged to develop a project that harnesses the power of experimental evolution, genomics, and the model yeast system Saccharomyces, to address fundamental questions in evolutionary biology.
Please apply through the official Stockholm University job platform no later than 30 April 2021: https://www.su.se/english/about-the-university/work-at-su/available-jobs?rmpage=job&rmjob=14698&rmlang=UK
10 March 2021
The lab has grown again! We welcome lab technician Pauline Caillault and MSc student Sofia de Beir into our team. Pauline has only been dissecting tetrads for a few weeks and is already threatening to beat my record of n spores/hour! Sofia has managed to move to Sweden from Portugal during these challenging times (and by that I don’t mean the snow outside, but also) and is now developing pipelines to analyze hundreds of (hybrid) genomes coming out of Ciaran’s 1000 generation evolution experiment for adaptive SNPs, structural variation, and aneuploidies. I am excited to have you both in the lab!
29 January 2021
The new year started great for us! Our paper Genomic evidence of an ancient East Asian divergence event in wild Saccharomyces cerevisiae is now published in Genome Biology and Evolution. This was a real team effort by several lab members, but especially the two lead authors Devin and Noah, who have worked really hard to analyze these interesting new genomes. Congratulations!

27 January 2021
This week, we had the awesome David Peris Navarro for a talk on yeast hybridization and a virtual lab visit. It was a lot of fun to compare notes (and genomes) with Peris, who had great input for the PhD students and postdocs in the lab. Thank you Peris for a great time, hope to meet you in person again soon!
5 December 2020
Rike got tenure!! I did some proper nerd baking for the occasion: a yeast population with standing genetic variation (the sprinkles are the SNPs) and a complete yeast haploid chromosome set with chocolate telomeres.



13 October 2020
We are welcoming a new group member! João has just started his PhD in population genetics. He is part of an interdisciplinary PhD programme that we share with Chun-Biu Li from the Department of Mathematics here at Stockholm University. Together with Chun and Devin, we want to better understand the impact of hybridization on adaption, using experimental evolution with yeast, fitness landscape theory and machine learning. Buckle up everyone, it’ll be a fun ride!
12 October 2020
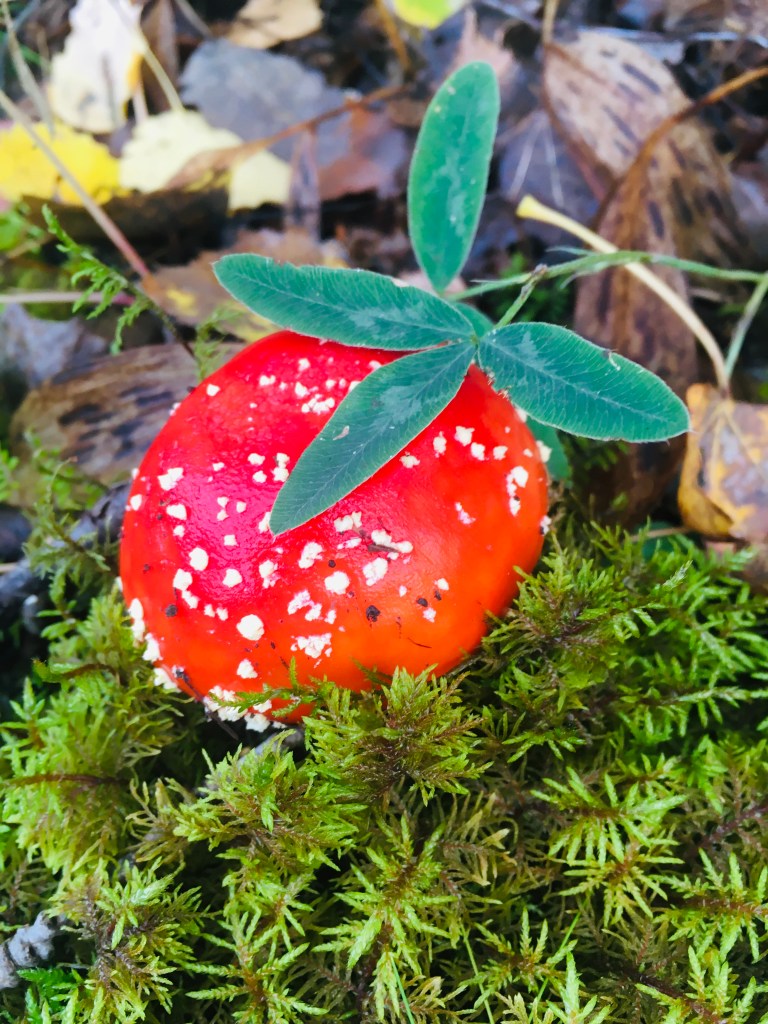
The lab went on a mushroom hunt in the woods. We got really lucky with the weather and Swedish forests are a magical place, full of trolls and fairies!








4 June 2020
We have a new preprint out! This is our first long read seq baby from the lab, and we are very excited about it. Devin and Noah have done an amazing job analyzing and meaningfully interpreting the vast structural genomic variation we found in seven divergent S. cerevisiae strains, isolated from wild habitats in China and Taiwan. We found large inversions and translocations (and maybe even a chromosomal fusion!) that don’t exist in other yeast genomes. Our phylogenetic comparisons suggest that some of these strains are the closest living relatives of the ancestors of all Baker’s yeast, from which they diverged before they went on and colonized the rest of the world. We hope these genomes will provide a nice testing field for other people interested in genome evolution!


11 May 2020
We are welcoming a new MSc student to the lab, Erik Zhivkoplias! Erik is working (together with Alex Rêgo) on how well different software tools perform to estimate selection coefficients, and how comparable the results of the different methods are. Erik got his hands on genomic time series data sets from very different model systems: yeast, seed beetles and Timema stick insects, so the results should be exciting! Luckily, Erik can work entirely remotely, which makes things much easier these days…
16 March 2020
Crazy times but something great has happened in the lab that I need to announce! Dragan Stajic (formerly IGC in Portugal) has joined us as a postdoc. Check out Dragan’s amazing work on epigenetic gene silencing with yeast here.
Dragan has come to our lab to work on the evolution of antibiotic resistance and the molecular basis of phenotypic switching, a mechanism by which one genotype produces two distinct phenotypes, which may help adaptation to complex environment.
Welcome, Dragan, we are excited to work with you!
7 March 2020
BIG NEWS! We have a PhD position available in “Understanding the Dynamics of Adaptation using Experimental Evolution with Yeast, Fitness Landscape Theory, and Machine Learning”. The position is part of a paired PhD programme between the Math Department and us, so supervision, lab meetings and seminars will be shared between departments to foster collaboration between biology, machine learning and statistical modelling.
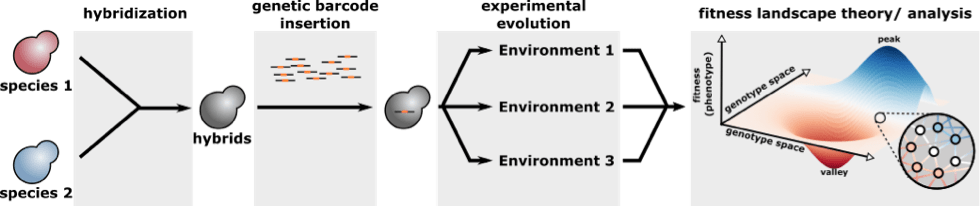
A full project description will follow soon but topics include 1) the dynamics of adaptation to changing environments with and without hybridization, 2) the construction of empirical fitness landscapes using DNA barcodes to track the evolution of lineages over time, 3) understanding the genetic architecture of (hybrid) fitness using machine learning and statistical modelling.
See Twitter and evoldir for the full advertisement of this position. All applications must go through the Stockholm University job portal: Follow this link to apply. Deadline for application is April 23.
Apply, apply!
4. February 2020
Winter is flying by and the light is back in Stockholm! We are proud to announce that PhD student Ciaran Gilchrist has successfully defended his half time thesis (Licentiate in Sweden) on Hybridisation, Aneuploidy and Adaptation to Environmental Stress in Baker’s Yeast. Some of Ciaran’s work is published here. We had the honour of hosting Niclas Backström (Uppsala University) as Ciaran’s external examiner who has provided much appreciated advice and input.
Also, I am happy to now officially be Docent at Stockholm University, a qualification similar to a habilitation in other European countries. Yay!
Finally, we are thrilled to see our very own Darwin-inspired yeast artwork on the cover of the January issue of Molecular Biology and Evolution! Drawing by me, photo by the talented Alessandro Devigili!

11. December 2019
Yesterday, we had the great pleasure to host Claudia Bank from the Gulbenkian Institute. Claudia gave a great talk on the the distribution of fitness effects of yeast mutations in different stressful environments, a topic close to our heart of course. It was great fun to chat about epistasis with Claudia, as always. Here is to next time, perhaps in a less cold and snowy town!
5. December 2019
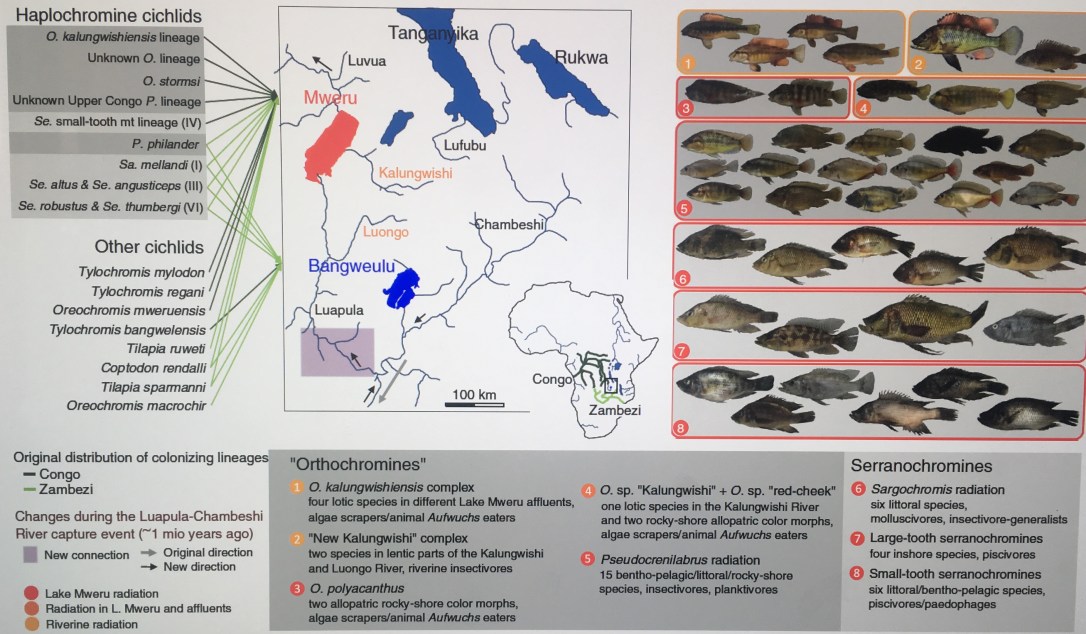
New paper out! See here what happens when African cichlid fish colonize a lake, find empty niche space, and hybridize! We found not just one, but 4 beautiful new adaptive radiations in Lake Mweru (on the Zambia/Kongo border).
https://www.nature.com/articles/s41467-019-13278-z

11. November 2019
Today Dragan Stajic from the Gulbenkian Institute in Lisbon visited the lab and told us about his very elegant experimental work with yeast on epigenetic gene silencing, the population genetic dynamics and interactions between epigenetic and genetic changes, how these become heritable and useful for adaptive evolution, and how closely intertwined they are with changes in the environment! Thanks for your visit, Dragan, looking forward to working with you!

6. November 2019
October was a whirlwind and it looks like things won’t slow down in November! We just had Walter Pfliegler visit the lab and tell us about his fascinating work on the micro- and macroevolutionary infection dynamics of pathogenic Saccharomyces in humans. Walter has a lot of exciting data and stories, look out for his publications! We are excited about our future collaborations with Walter.
Two weeks ago we had our yearly departmental retreat (aptly named “The Bloodbath” for historical reasons) where mainly students and postdocs present their projects to get feedback from faculty and invited speakers, this year featuring speciation genetics experts Kerstin Johannesson (Littorina snails) and Jochen Wolf (crows). The meeting takes place in our beautiful field station Tovetorp (we got to cuddle those fallow deer babies!). This year, Devin, Zebin, Ciaran and Alex from the lab presented their work and inspired us all with their yeasty genius.
8. October 2019
We had the absolute please to host Patrik Nosil (CEFE, Montpellier) this week. It was great to talk to Patrik about fitness landscapes, epistasis, speciation, stick insects, and experimental evolution with yeast. Such an inspiring and fun visit, thanks Patrik!
7. October 2019
We have budded again this week! We are welcoming a new lab member, Noah Gettle! Noah has just finished his PhD in Mike Travisano’s lab and is now off to new yeast adventures in our lab. Welcome, Noah! Looking forward to working with you.
14. September 2019
New paper out in Molecular Biology and Evolution led by Devin Bendixsen and Zebin Zhang! Read here how recombination puzzle-pieces together F2 hybrid genomes from vastly divergent species and still makes a fit and fertile individual!

2. August 2019
Whohoo! Ciaran’s opinion paper on hybridization and aneuploidy in yeast, and how both might contribute to adaptation, is now online and open access in Yeast!
https://onlinelibrary.wiley.com/doi/full/10.1002/yea.3427

31. July 2019
In other news, I received this plant for my birthday! Thanks lab, you are as sweet as this bonsai pineapple (will hopefully be one day if I don’t kill it).

30. July 2019
With a bit of a delay, welcome to undergrad student Viktoria Köppa! Viktoria is doing an internship in our lab, working on fine-tuning different stressful environments for our evolution experiments.
28 May 2019
We are thinking about rebranding the lab… wonder if that’s ok with SU?

17 May 2019
The Knut and Alice Wallenberg Foundation published a little blurb about our research on their website. Read here. And no, I do not spend my days gazing at dissected tetrads under the fume hood. Especially not without a lab coat! 🙂
7 May 2019
Our lab won the departmental photo competition, wohoo! Here is the winner (idea and photo by Alessandro Devigili, streaking by Rike). We call it “One tree to rule them all”! 🙂

12. February 2019
Welcome to Julie Grosse Sommer who is visiting us from Maastricht University this spring semester. She is working on her BSc thesis and competing yeast hybrids against their parents in the lab!
9. January 2019
We are looking for enthusiastic Master’s students who want to work with experimental evolution and Saccharomyces budding yeast in beautiful Stockholm. We have exciting projects available, e.g. on temperature-dependent fitness effects of outcrossing!
7. January 2019
Happy new year everyone, and congratulations to my PhD student Ciaran Gilchrist for winning a travel grant from the Alice och Lars Siléns Fond! Well done, Ciaran!
11. December 2018
Congrats to postdoc Devin Bendixsen for winning a prestigious postdoctoral fellowship for incoming researchers from the Wenner Gren Foundation! Really looking forward to continue working on hybrid fitness landscapes with you, Devin!
11. December 2018
We had our very first guest speaker over today, Dr. David Berger from Uppsala University! David gave a fascinating talked packed with beautiful data on the condition-dependence of mutation rates and the effect of temperature on the strength of selection. Here’s hoping to a fun and productive collaboration between our labs!
9. December 2018
I went to a fancy reception in honour of the 2018 Nobel Laureates today, in the stately Nordiska Museet of Stockholm. I only got a glimpse of Frances Arnold, but the champagne and the pickled herring appetizers were delish!
9. November 2018
We finally have our own lab Twitter account, many thanks to Alex Rêgo! https://twitter.com/StelkensLab
16. October 2018
We have just uploaded ideas for interesting Master’s research projects in the lab. Have a look at the Stockholm University Master Project page for more information. If you are interested, drop me an email!
3. October 2018
We are excited to announce that we are looking for 2 more postdocs to join the Stelkens lab! For more information see this ad on evoldir.
1. October 2018
A warm Välkommen till Sverige and the Stelkens Lab to our new lab members Devin Bendixsen and Alexandre Rêgo! Devin just finished his PhD on some very cool and innovative empirical fitness landscapes at Boise State University in Idaho, and Alex just graduated with an MSc from Utah State University on rapid adaptation in the seed beetle. From one charismatic model system to the next, Alex! Ironically, even though Devin and Alex moved around the world for their new adventures in Europe, they are now reunited in the same office.

